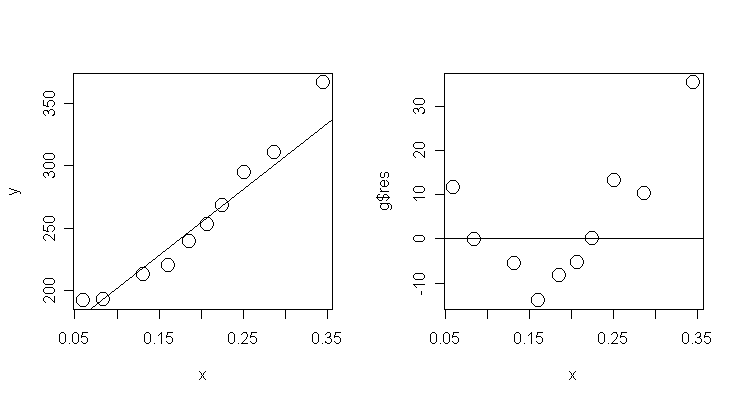Testing
for lack of fit
(Reading:
Faraway(2005, 1st edition), 6.3)
Variance known
-
Using
the same data as in
the weighted least
squares lab, we are going to test whether the model fitted in the previous
lab is adequate.
-
Because of
the way the weights are defined, i.e., wi=1/var(yi), the known
variance is implicitly equal to one (why?).
Let's first fit the weighted least square model.
> p <- read.table("phy.data", header=T)
# read the data into R
> p # take a look of the data
momentum
energy crossx sd
1 4
0.345 367 17
2 6
0.287 311 9
3 8
0.251 295 9
...deleted...
10 150
0.060 192 5
> x <- p[,2]; y <- p[,3]; w <-
1/p[,4]^2
# extract variables and calculate
weights
> g <- lm(y~x,weights=w) # fit a simple regression model with w
as weights
> summary(g)
# take a look of the fitted model
Call:
lm(formula = y ~
x, weights = w)
Residuals:
Min 1Q Median 3Q Max
-2.323e+00
-8.842e-01 1.266e-06 1.390e+00 2.335e+00
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept)
148.473 8.079 18.38 7.91e-08 ***
x
530.835 47.550 11.16 3.71e-06 ***
---
Signif. codes: 0
'***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard
error: 1.657 on 8 degrees of freedom
Multiple
R-Squared: 0.9397, Adjusted R-squared: 0.9321
F-statistic:
124.6 on 1 and 8 DF, p-value: 3.710e-06
-
Look at the high R2 and the
very small p-value for overall F-test
-
Q: do you think the model is a good fit?
-
Now
plot the data and the fitted regression line:
>
par(mfrow=c(1,2)) #
split the graphic window into two
> plot(x, y, cex=2) # draw the scatter plot of energy and
crossx
> abline(g$coef) # add the fitted line
Now, let's also plot the residuals against crossx.
> plot(x, g$res, cex=2); abline(0, 0)
# draw scatter plot of residuals and crossx and add y=0
line
-
Let's perform the test for lack of fit. Compute
the test statistic and its p-value:
> (8*1.657^2)/1 # test statistics = (n-p)*σ-hat2/σ2
> 1-pchisq(21.96519, 8) # p-value of the test, the "pchisq"
command returns the cdf value of a chi-square distribution
-
Because the p-value is small, we
conclude that there is a lack of fit.
-
Notice that although the original linear
model had an R2 of 94% and a very small overall F p-value, this in
itself is not enough to conclude that this particular model fits.
Let us try to
add a quadratic term, as suggested by the residual plot above, to the model and test again:
> g1 <- lm(y ~ x + I(x^2), weights=w) # fit a 2nd-order polynomial model
>
summary(g1, cor=T) # take a look of the new fitted model
Call:
lm(formula = y ~
x + I(x^2), weights = w)
Residuals:
Min
1Q Median 3Q Max
-0.89928
-0.43508 0.01374 0.37999 1.14238
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept)
183.8305 6.4591 28.461 1.7e-08 ***
x
0.9709 85.3688 0.011 0.991243
I(x^2)
1597.5047 250.5869 6.375 0.000376 ***
---
Signif. codes: 0
'***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard
error: 0.6788 on 7 degrees of freedom
Multiple
R-Squared: 0.9911, Adjusted R-squared: 0.9886
F-statistic:
391.4 on 2 and 7 DF, p-value: 6.554e-08
Correlation of
Coefficients:
(Intercept) x
x
-0.94
I(x^2)
0.86 -0.97
-
The quadratic term is significant.
-
The linear term
becomes insignificant because of the strong collinearity between linear and
quadratic terms.
-
Let's perform the test for lack of fit on the new
model.
> (7*0.6788^2)/1 # calculate test statistic
> 1-pchisq(3.225386, 7) # calculate p-value
-
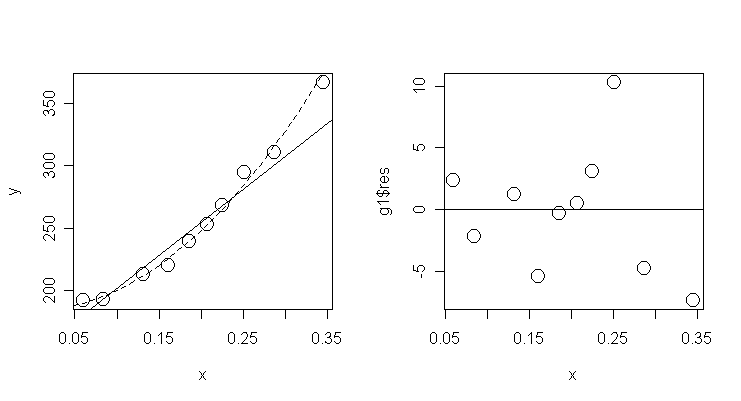
Let's plot the fits of the two models and a
residual plot under the new model:
> plot(x, y, cex=2) # draw the scatter plot of energy and
crossx
> abline(g$coef) # add the fitted line of model g
> xm <- seq(0.05, 0.35, 0.01); lines(xm, g1$coef[1]+g1$coef[2]*xm+g1$coef[3]*xm^2, lty=2) # add
the fitted line of model g1
> plot(x, g1$res, cex=2); abline(0, 0)
# a residual plot under model g1
Variance unknown
-
The
data
for this example consist of thirteen specimens of 90/10 Cu-Ni alloys with
varying iron content in percent.
-
The specimens were submerged in sear water for
60 days and the weight loss due to corrosion was recorded in units of milligrams
per square decimeter per day.
> corrosion <-
read.table("corrosion.data") # read the data into R
> corrosion
# take a look of the data
Fe loss
1 0.01 127.6
2 0.48 124.0
3 0.71 110.8
4 0.95 103.9
5 1.19 101.5
6 0.01 130.1
7 0.48 122.0
8 1.44 92.3
9 0.71 113.1
10 1.96 83.7
11 0.01 128.0
12 1.44 91.4
13 1.96 86.2
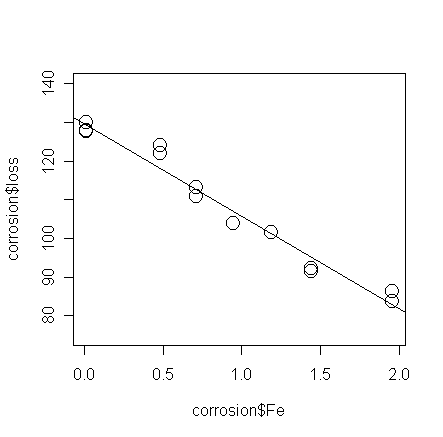
> g <- lm(loss ~ Fe, data=corrosion) # Fit
a simple linear model with loss as response and Fe as predictor
>
summary(g) # take a look of the fitted model
Call:
lm(formula = loss
~ Fe, data = corrosion)
Residuals:
Min 1Q
Median 3Q Max
-3.7980 -1.9464
0.2971 0.9924 5.7429
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept)
129.787 1.403 92.52 < 2e-16 ***
Fe
-24.020 1.280 -18.77 1.06e-09 ***
---
Signif. codes: 0
'***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard
error: 3.058 on 11 degrees of freedom
Multiple
R-Squared: 0.9697, Adjusted R-squared: 0.967
F-statistic:
352.3 on 1 and 11 DF, p-value: 1.055e-09
To perform the test for lack of fit,
> ga <- lm(loss ~ factor(Fe),
data=corrosion) # fit the saturated model, the
"factor" command treats Fe as a qualitative predictor
> summary(ga)
# take a look of the fitted saturated model
Call:
lm(formula = loss
~ factor(Fe), data = corrosion)
Residuals:
Min 1Q Median 3Q Max
-1.250e+00
-9.667e-01 9.991e-17 1.000e+00 1.533e+00
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept)
128.567 0.809 158.914 4.19e-12 ***
factor(Fe)0.48
-5.567 1.279 -4.352 0.00481 **
factor(Fe)0.71
-16.617 1.279 -12.990 1.28e-05 ***
factor(Fe)0.95
-24.667 1.618 -15.245 5.03e-06 ***
factor(Fe)1.19
-27.067 1.618 -16.728 2.91e-06 ***
factor(Fe)1.44
-36.717 1.279 -28.703 1.18e-07 ***
factor(Fe)1.96
-43.617 1.279 -34.097 4.24e-08 ***
---
Signif. codes: 0
'***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard
error: 1.401 on 6 degrees of freedom
Multiple
R-Squared: 0.9965, Adjusted R-squared: 0.9931
F-statistic:
287.3 on 6 and 6 DF, p-value: 4.152e-07
Notice that
-
the saturated model has 7 parameters
(including a constant term), which is equal to the number of distinct values of
Fe.
-
The residual standard error, 1.401, is a pure error estimate of true
σ.
-
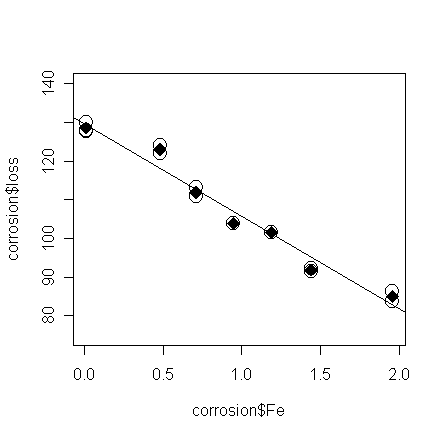
The
fitted values of the saturated model are the means in each group - put these on the plot:
> points(corrosion$Fe, ga$fit, pch=18, cex=2) # add the means in each group to the plot
-
Here, let's calculate the SS and degrees of
freedom for the pure error "by hand":
> sum(ga$res^2) # calculate SSp.e.
>
ga$df # the dfp.e.
> sum(g$res^2) # the RSS
calculated from the model g
We can put these values into the ANOVA table to
calculate test statistic:
| |
d.f. |
SS |
MS |
F |
|
Residual |
n-p
(11) |
RSS
(102.8502) |
|
|
|
Lack of fit |
n-p-dfp.e.
(5) |
RSS-SSp.e. |
(RSS-SSp.e.)/(n-p-dfp.e.
) |
ratio of MS |
|
Pure error |
dfp.e.
(6) |
SSp.e.
(11.78167) |
(SSp.e.)/(dfp.e.) |
|
And, now the test statistic:
> ((102.8502-11.78167)/(11-6))/((11.78167/6)) # test statistic for lack of fit
> 1-pf(9.275615, 5, 6) # the p-value
-
A
slightly quicker way to get the same result:
> anova(g, ga)
# compute the ANOVA table for models g
and ga
Analysis of
Variance Table
Model 1: loss ~
Fe
Model 2: loss ~
factor(Fe)
Res.Df RSS
Df Sum of Sq F Pr(>F)
1 11
102.850
2 6 11.782
5 91.069 9.2756 0.008623 **
---
Signif. codes: 0
'***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Let's
try to add more higher order polynomial terms one-by-one until the model pass
the test for lack of fit:
> g1 <- lm(loss~Fe+I(Fe^2), data=corrosion)
# fit a 2nd-order polynomial model
>
anova(g1, ga) # compute the
ANOVA table for models g1 and ga
Analysis of
Variance Table
Model 1: loss ~
Fe + I(Fe^2)
Model 2: loss ~
factor(Fe)
Res.Df RSS
Df Sum of Sq F Pr(>F)
1 10
100.086
2 6 11.782
4 88.305 11.243 0.005949 **
---
Signif. codes: 0
'***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
> g2 <-
lm(loss~Fe+I(Fe^2)+I(Fe^3), data=corrosion) # fit a
3rd-order polynomial model
> anova(g2, ga)
# compute the ANOVA table for models g2
and ga
Analysis of
Variance Table
Model 1: loss ~
Fe + I(Fe^2) + I(Fe^3)
Model 2: loss ~
factor(Fe)
Res.Df RSS
Df Sum of Sq F Pr(>F)
1 9
45.051
2 6 11.782
3 33.269 5.6476 0.03506 *
---
Signif. codes: 0
'***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
> g3 <-
lm(loss~Fe+I(Fe^2)+I(Fe^3)+I(Fe^4), data=corrosion) #
fit a 4th-order polynomial model
>
anova(g3, ga) # compute the
ANOVA table for models g3 and ga
Analysis of
Variance Table
Model 1: loss ~
Fe + I(Fe^2) + I(Fe^3) + I(Fe^4)
Model 2: loss ~
factor(Fe)
Res.Df RSS
Df Sum of Sq F Pr(>F)
1 8
34.975
2 6 11.782
2 23.194 5.9059 0.03822 *
---
Signif. codes: 0
'***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
> g4 <-
lm(loss~Fe+I(Fe^2)+I(Fe^3)+I(Fe^4)+I(Fe^5), data=corrosion)
# fit a 5th-order polynomial model
>
anova(g4, ga) # compute the
ANOVA table for models g4 and ga
Analysis of
Variance Table
Model 1: loss ~
Fe + I(Fe^2) + I(Fe^3) + I(Fe^4) + I(Fe^5)
Model 2: loss ~
factor(Fe)
Res.Df RSS
Df Sum of Sq F Pr(>F)
1 7
14.465
2 6 11.782
1 2.683 1.3663 0.2868
Finally,
the 5th-order polynomial model passes the test for lack of fit.
-
This model is very
close to the saturated model (the 6th-order polynomial model is a saturated model,
why? Hint. count the number of
parameters).
-
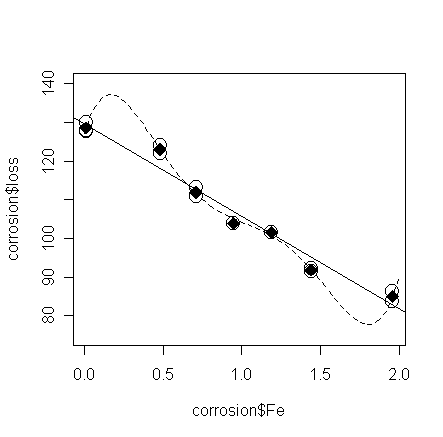
Let's plot the fitted line of the 5-th order polynomial model and compare
it with the straight line model:
> grid <- seq(0, 2, length=50);
lines(grid, predict(g4,data.frame(Fe=grid)), lty=2) # add the fitted line of the 5-th order
model